CHEMICAL BIOLOGY OF NUCLEIC ACIDS
Our research group is interested in the chemical synthesis, biochemical properties, and molecular behaviour of nucleic acids and their analogues. We are particularly interested in developing new methods for the synthesis of nucleoside building blocks and RNA, including modified siRNA, ASOs,crRNA, branched RNA, and lariat RNA. We also devote an enormous effort to the study of nucleic acid structure.
The main methods used in these investigations are solution and solid-phase synthesis; molecular biology techniques (gene silencing via RNAi, antisense, PCR, etc), high resolution NMR, UV and circular dichroism; and molecular modeling.
Click on a research theme below to learn more.
Antisense Oligonucleotides & Gene Silencing
Frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS) are neurodegenerative diseases often caused by repeat expansion within the non-coding region of the chromosome 9 open reading frame 72 (C9orf72) gene. Both DNA strands are transcribed into long RNA which aggregate as foci in the cell nuclei and are translated into dipeptide repeat proteins (DPRs) in the cytoplasm. Accumulation of the foci and DPRs is toxic to cells within the central nervous system and leads to progressive deterioration in the patient’s health. Antisense oligonucleotides (ASOs) are an emerging class of therapeutics for genetic diseases that function through modulation of different components related to the disease phenotype. ASOs can be programmed to hybridize to complementary target RNA and either sterically block ribosomal translation or recruit RNase H endonucleases for degradation. Both mechanisms lead to gene knockdown and can be used to reduce the toxic phenotypic effects of the repeat expansion found in patients. We designed a series of ASOs targeting the G- and C-rich repeats of C9orf72. These studies provide insight into the genetic cause of these diseases and the design of therapeutic oligonucleotides to treat this disease (H. Barber).
Duchenne muscular dystrophy (DMD) is a neuromuscular disease caused by mutations in the dystrophin gene. The deletion of exons leads to a frameshift in the mRNA and the production of a truncated, non-functional form of the dystrophin protein. Exons 51 and 53 can be skipped and the recovery of a readable frame leads to a shorter but functional form of the protein. This can be achieved with ASOs. Approved exon skipping ASOs on the market are based on phosphorodiamate morpholino oligomers (PMO) and phosphorothioate (PS) 2’-OMe modifications; however, their approval is controversial, and observed levels of dystrophin increase were minor with unclear therapeutic relevance. We designed novel modification patterns that produce unprecedented skipping levels for exon 53 in vitro and optimistic results in vivo. For exon 51, an issue arose with the apparition of unwanted cryptic splicing products. To recover the correct splicing product and increase cellular uptake, we are pursuing ASOs containing new patterns as well as increasing muscular uptake for exon 53 with oligonucleotide conjugates (E. Groenwold).
Nucleic Acid Structure
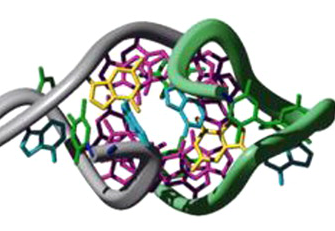
Z-forming sequences play a pivotal role in regulating various biological processes, including genome stability and immune responses, making them a focus of intense research. Despite their biological significance, the in vitro stabilization of Z-DNA and Z-RNA has traditionally required extreme conditions such as high salt concentrations or specific solutes to overcome their inherent instability. Our recent studies have demonstrated that 2’-fluorinated nucleosides, specifically 2’-F-ana (2’-fluoro-arabinose) and 2’-ribo-F sugars, showed enhanced propensity of nucleic acids to adopt and maintain the Z-conformation under physiological conditions, expanding their potential for biological and therapeutic applications. (H.Kaur)
Single-stranded nucleic acid aptamers fold into defined three-dimensional conformations like G-quadraplex, hairpins, loops etc.; allowing them to bind their targets with high affinity and specificity. While aptamers offer distinct advantages over antibodies, being non-immunogenic, non-toxic, and highly stable, their clinical translation has been limited by poor nuclease stability and bioavailability. To overcome these challenges, we employ sugar-modified nucleosides such as 2′-fluoro-arabinonucleic acid (2′F-ANA) nucleosides at specific positions. These modifications refine the physicochemical properties of the aptamer, especially structural stability and nuclease resistance, while ensuring its spatial conformation remains intact, thereby enhancing its therapeutic efficacy. (H.Kaur)
Branched and Long RNA
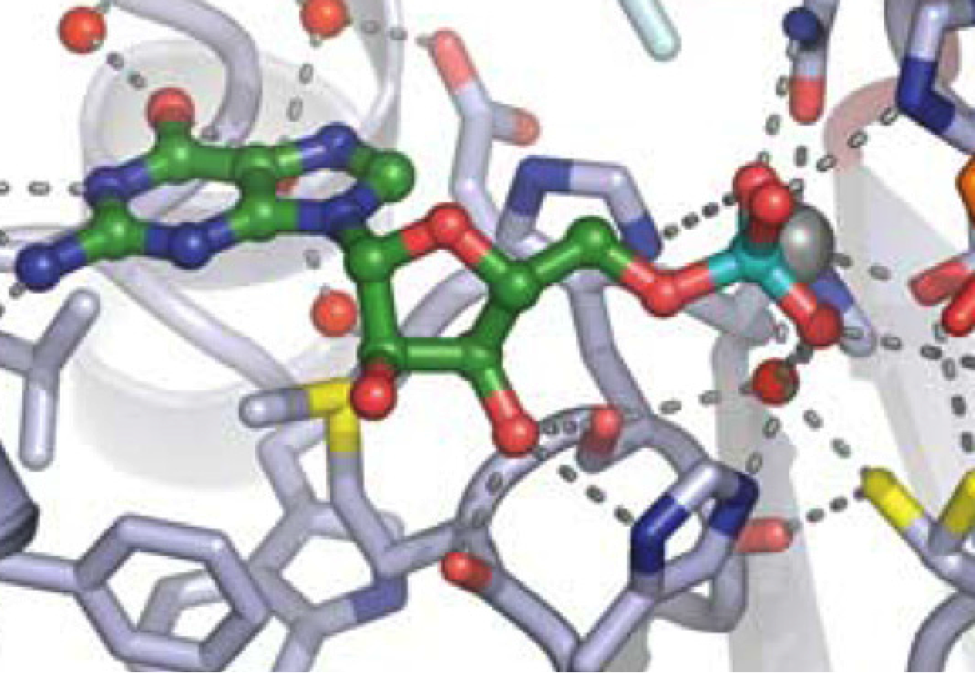
Intron lariats are naturally occurring branched RNA intermediates that possess an unusual 2’-5’ phosphodiester bond originated from the eukaryotic splicing of the pre-mRNA. The RNA lariat debranching enzyme (Dbr1) is a eukaryotic metallophosphodiesterase that can hydrolyze such 2’-5’ phosphodiester linkage. The accumulation of branched RNAs caused by Dbr1 deficiency is likely to associate with suppressed innate immune response in humans. This happens by inhibiting one or more of the cellular antiviral dsRNA sensors including TLR3, RIG-I, MDA5, PKR, OAS1, OAS2, and OAS3. Studying the mechanism of the blurring effect of the lariat RNA on the activity of the known dsRNA sensors is essential to understand the susceptibility of viral infection to individuals with Dbr1 deficiency. We designed a series of synthetic 2’-branched RNAs (bRNAs) analogues to mimic the intronic RNA lariats. The inhibitory activity of dsRNA sensors by our synthetic branched RNA analogues are examined to support the notion that viral infection susceptibility triggered by Dbr1 deficiency is due to RNA lariat accumulation (Z. Lyu).
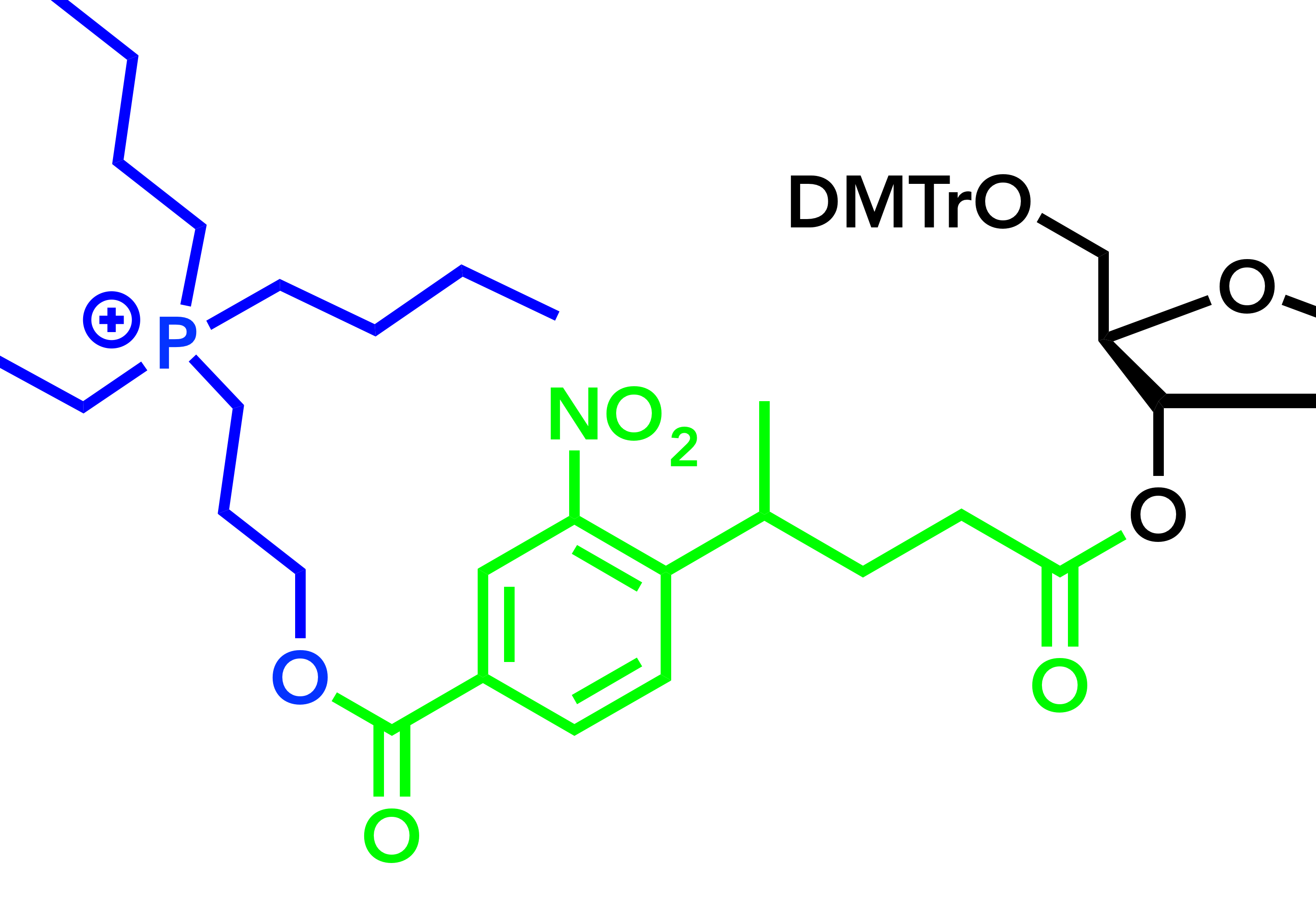
The onset of CRISPR-Cas9 technology created a need for synthetic single-guide RNAs (sgRNA), which are 99 nucleotides long, and the necessary rapid iterative screening efforts require a high synthetic throughput. Current solid-phase technology typically applies 2´-O-tert-butyldimethylsilyl (TBDMS) phosphoramidites, which can produce 20-100mer oligonucleotides, but require 10-15 min coupling times for optimal performance. Considering 100 couplings per oligonucleotide, this exposes the growing sequence to prolonged acidic conditions and the overall synthesis time exceeds 24h. Moreover, TBDMS deprotection sequence requires a fluoride treatment, which must follow the standard base deprotection, and must be eliminated before clinical application. Phosphoramidites featuring 2´-O-acetal levulinic ester (ALE) groups can efficiently produce RNA on both solid support and a photolithographic chip. We investigate the use of ALE chemistry to produce sgRNA, and identified deprotection conditions that improve the synthesis speed, quality, and quantity of sgRNA and other long RNAs. (Z. Lyu).
Green Approaches to Oligonucleotide Synthesis
Recent advances in nucleic acid therapeutics have drastically increased the demand for synthetic oligonucleotides (ONs). While current solid-phase oligonucleotide synthesis (SPOS) protocols are highly efficient and automated, they require huge amounts of solvent and a high excess of reagents. Mechanochemistry has recently emerged as a new tool to explore novel reactions. Mechanical action, such as vibration ball milling (VBM), provides the energy and mixing for chemical reactions to take place, often completely in the absence of solvent. A recent analysis of several ONs in different stages of development examined the process mass intensity (PMI) for their synthesis and demonstrated the incredible amount of waste generated during ON synthesis. They further subdivided the analysis to PMI contribution by material type, process stage, and synthesis stage. The contribution of wash solvents and reaction solvents consisting of organic solvents is of note. Although a large contribution to the PMI is from water during purification, there is a need to address the significant contribution from wash and reaction solvents. Our work seeks to alleviate these issues by synthesizing oligonucleotides using mechanochemistry using little to no solvent (J. Thorpe).
Biologics have become a powerful tool in combating diseases inaccessible to small molecules. At the forefront of this new wave in drug development are oligonucleotide-based therapies. CRISPR technologies, antisense oligonucleotides, and siRNA are powerful nucleotide-based tools which can modify the expression, transcription, and fundamental structure of our genetic code. These technologies have great potential in the amelioration of genetic conditions, with nineteen approved nucleic acid therapeutics on the market, and many more in the clinic. However, the synthesis of these oligonucleotides has presented a hurdle, and while the advent of SPOS has made development and commercialization of oligonucleotides feasible, it is insufficient in both environmental and economic terms. For this reason, it is vital to investigate new synthetic strategies. We demonstrate the utility of resonant acoustic mixing (RAM) in the synthesis of oligonucleotides utilizing phosphoramidite chemistry. RAM offers a mechanochemical medium which is both gentler and more amenable to scale-up reactions than other mechanochemical methods. We have developed solvent and liquid free RAM phosphoramidite protocols which give high yields, offering a green alternative to traditional solid phase synthesis (J. Marlyn).
CRISPR-Cas Systems & Gene Editing
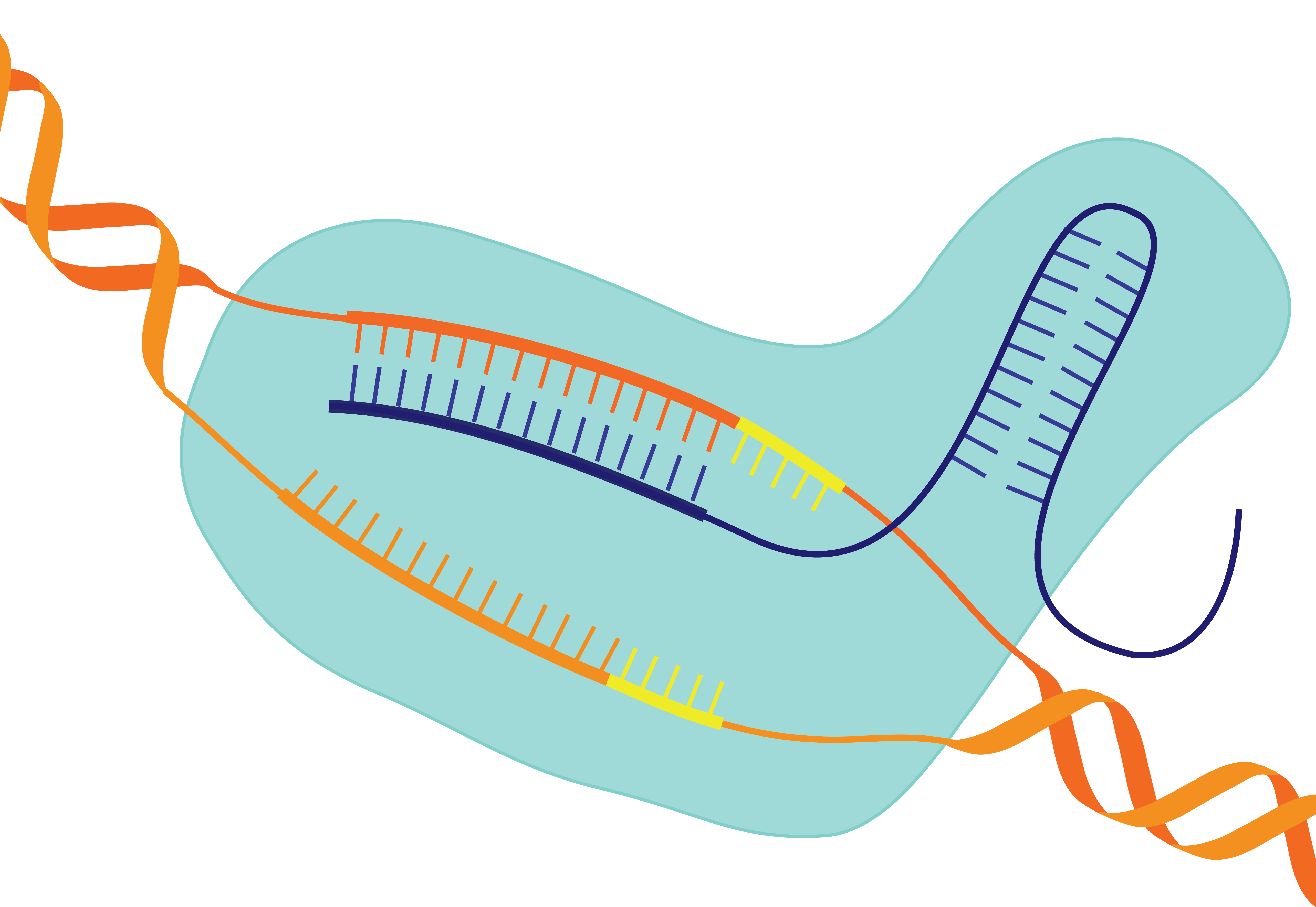
Clustered regularly interspaced short palindromic repeats (CRISPR) technology, originally discovered as an innate bacterial immune system against foreign nucleic acids, has been harnessed in recent years as a gene editing tool for use in mammalian cells. Often referred to as genetic scissors, CRISPR-Cas systems consist of an RNA-guided endonuclease composed of a CRISPR-associated (Cas) effector protein and a CRISPR RNA (crRNA). To improve the drug-like properties of the crRNA so that CRISPR technology can be used to treat genetic diseases, it must be chemically modified through the incorporation of nucleotide analogues. While many other therapeutic oligonucleotides have been readily modified for clinical use, modification of the crRNA remains challenging due to its intricate nucleic acid-protein interactions. We use a structure-guided approach is used to systematically incorporate chemically modified nucleotides into the sequence without negatively impacting gene editing efficiency (H. Barber & M. Cunningham).
CRISPR-based therapeutics stand to revolutionize treatment for genetic illnesses. To fully implement CRISPR-based therapeutics, it may be necessary to develop kill-switch inhibitors that can halt activity on demand. Existing methods of inhibiting CRISPR include naturally occurring anti-CRISPR proteins, which exhibit high binding affinities but present delivery challenges, and some small molecules, which are easily administered but display low binding affinities. Rationally designed small modified nucleic acid-based inhibitors (SNuBs) display a large degree of specificity for a certain target, as well as high binding affinities due to Watson-Crick-Franklin base-pairing, providing an easier-to-deliver alternative to anti-CRISPR proteins. However, both the design and synthesis of these SNuBs require optimization (M. Cunningham).
Synthesis of RNA Mimics
Our lab is dedicated to the design, synthesis, and biological evaluation of 2′, 4′, and 2′,4′-ribose-modified RNA analogues in the context of nucleic acid-based therapeutics. Chemical modification of RNA allows us to elucidate the roles of discrete functional groups within its structure and facilitate the study of RNA–ribonucleoprotein interactions. These modifications enhance binding affinity, improve nuclease resistance, and reduce immune activation, all of which are essential for transforming RNA into a viable therapeutic with superior pharmacokinetic and pharmacodynamic properties. Therapeutic applications of CRISPR require extensive chemical modifications like those used in clinically approved oligonucleotide therapeutics. However, certain regions of crRNA necessitate the presence of a 2′-OH group for optimal gene-editing efficiency, presenting a challenge for Cas9, Cas12a, and related enzymes. While key nucleotide analogues such as 2′-F and 2′-OMe enhance binding affinity, improve nuclease stability, and mitigate immune activation, they lack the crucial 2′-OH groups, limiting their utility at these critical positions in CRISPR-crRNA applications. To overcome this challenge, we are developing RNA mimics that preserve essential 2′-OH interactions necessary for effective Cas-mediated gene editing. (A. Arora & E. Cordova).